| Prev | ICM User's Guide 22.5 Sequence and Alignment Tutorial | Next |
[ Load PDB Structures | Load Sequences | Link Sequence to Structure | Pocket Sequence Conservation ]
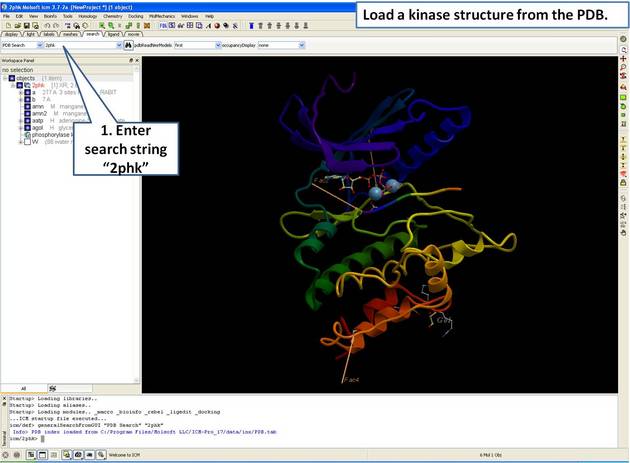
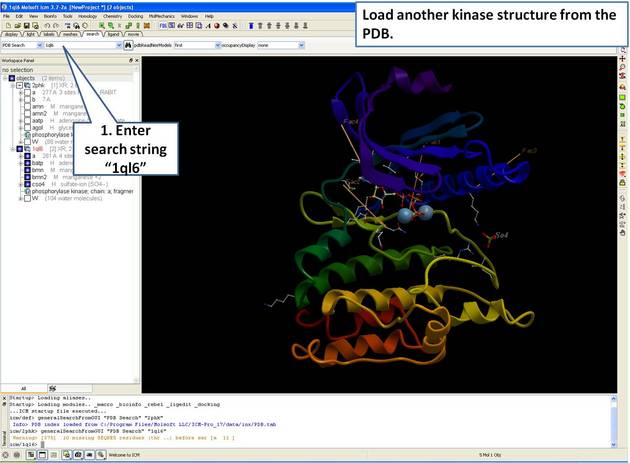
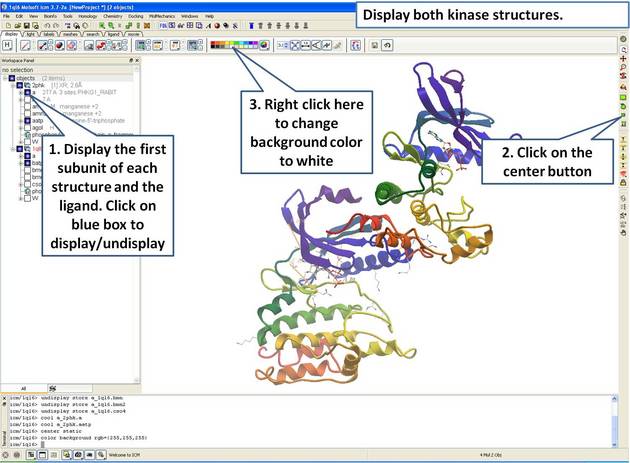
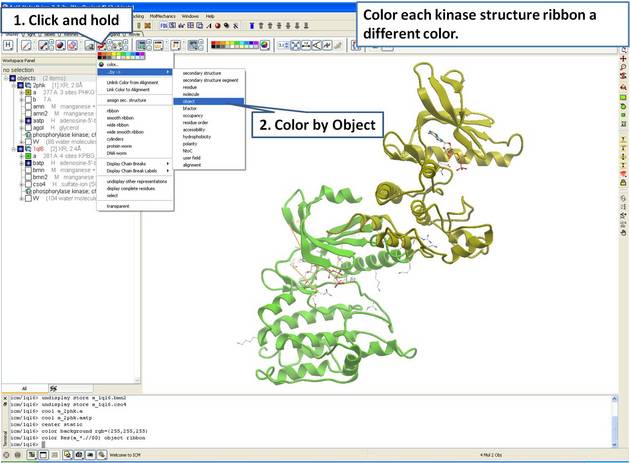
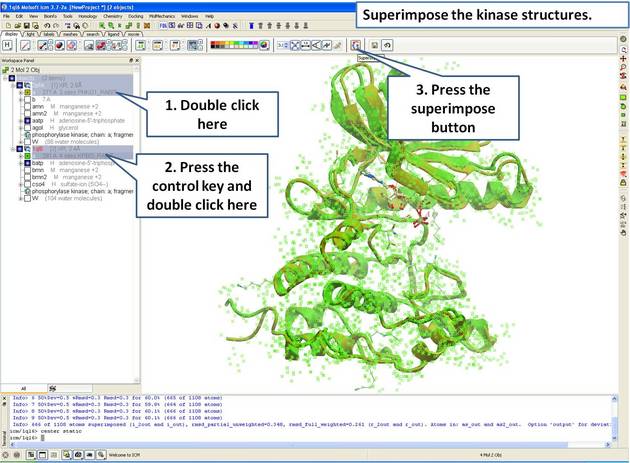
22.5.1 Load and Display Protein Kinase Structures |
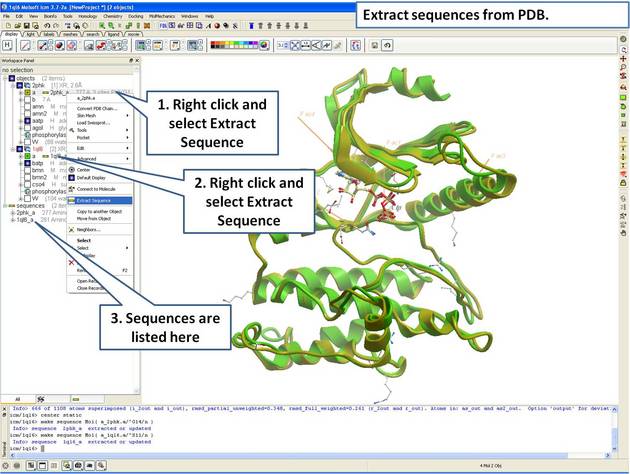
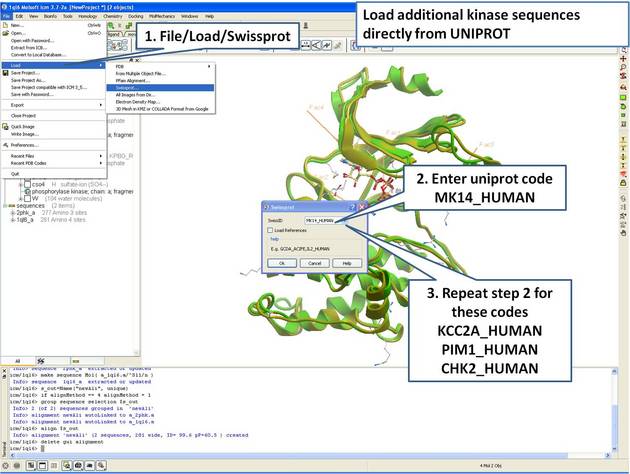
22.5.2 Extract Sequences from PDB Structures and Load New Sequences from UniProt |
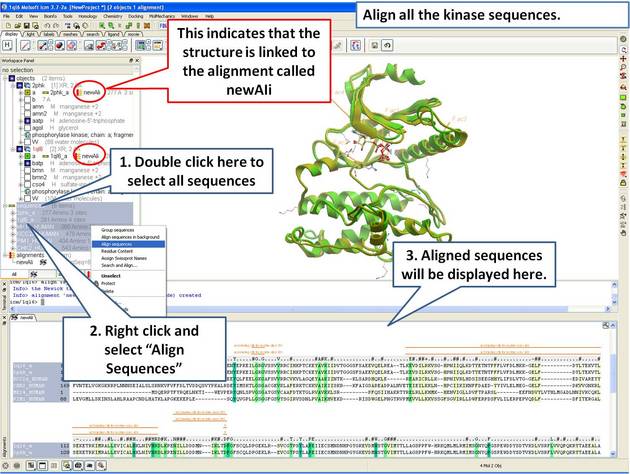
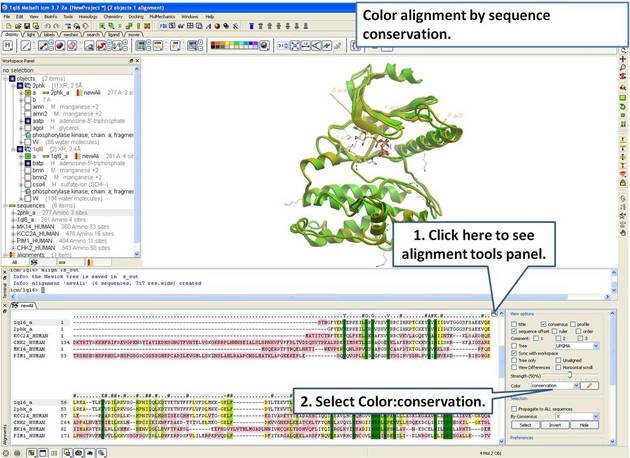
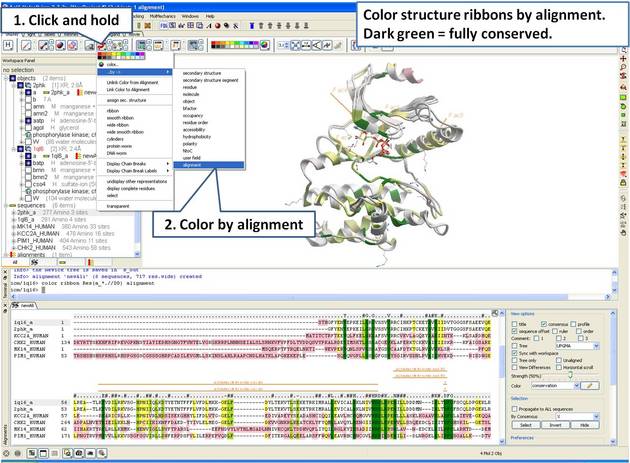
22.5.3 Linking Sequence Alignment to Structure |
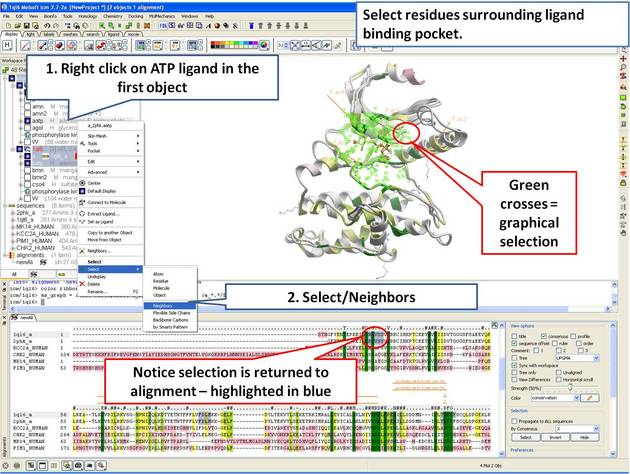
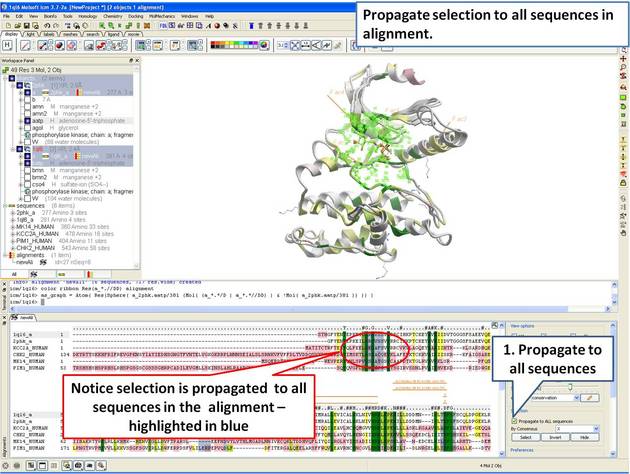
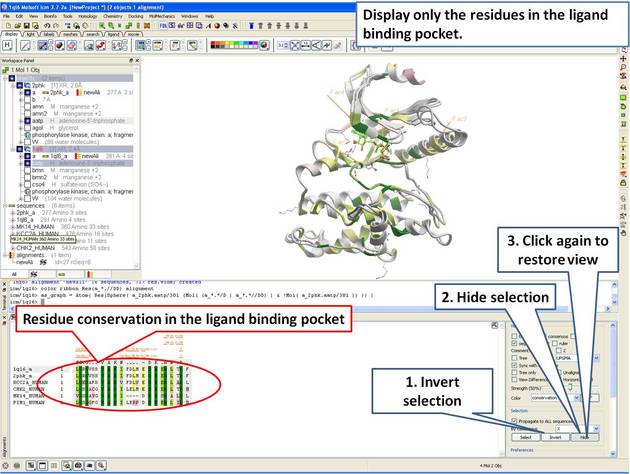
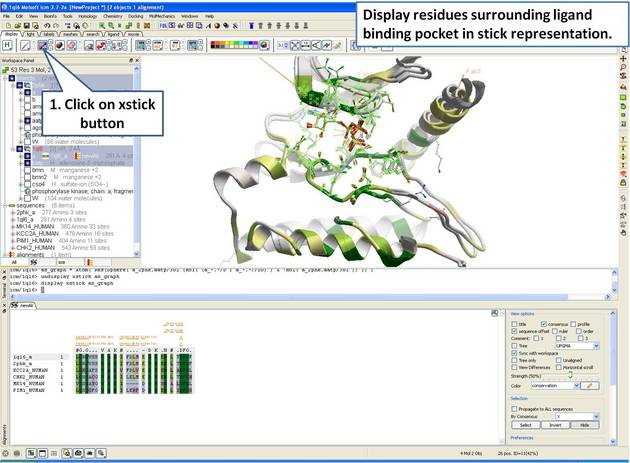
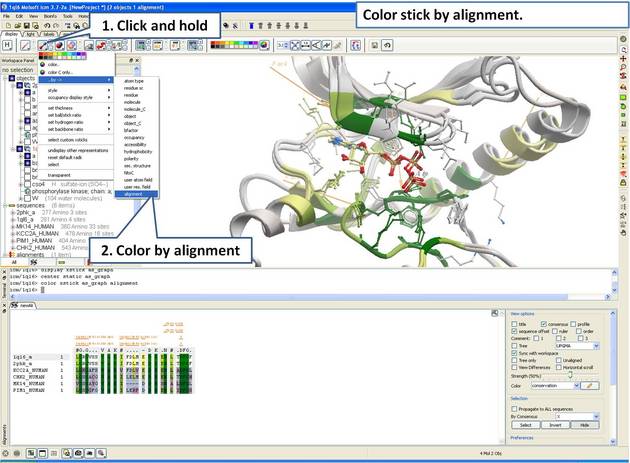
22.5.4 Identify Sequence Conservation in Ligand Binding Pocket |
| Prev Convert | Home Up | Next Ligand Binding Pocket Analysis |