| Prev | ICM User's Guide 22.8 Protein Preparation and Crystallographic Analysis Tutorial | Next |
[ PDB Preparation - Symmetry | PDB Preparation - Occupancy | PDB Preparation - Alternative Orientation | Biomolecule ]
| Note: Click Next (top right hand corner) to navigate through this chapter. Headings are listed on the left hand side (web version) or by clicking the Contents button on the left-hand-side of the help window in the graphical user interface. |
22.8.1 PDB Preparation - Symmetry |
Background When inspecting a ligand binding pocket it is important to check that the true pocket is formed by chains which are not explicitely present in a PDB entry. Therefore it is necesary to use Tools/X Ray/Crystallographic Neighbor to find all molecules/subunits or chains involved in the interaction with the ligand. Molecular objects and 3D density maps may contain information about crystallographic symmetry. It consists of the following parameters:
- Crystallographic group eg. P2121 that determine N (depends on a group) transformations for the atoms in the asymetric unit.
- Crystallographic cell parameters A, B, C, Alpha, Beta and Gamma
Example As an example let us look at Cycloldextrin glycosyltransferase (PDB Code: 1CDG). The problem with docking to this receptor is that the true pocket is formed by chains which are not explicitly present in the PDB entry. Site mb1 includes serine 382. This cannot be predicted just by looking at the structure. Therefore we need to identify symmetry related molecules to this protein.
- Use the PDB search tab to load the crystal structure 1cdg.
- Inspect the ligand binding pocket of maltose (mal)
- To identify if there are any other chains involved in the interaction with the ligand select the whole structure in the ICM Workspace.
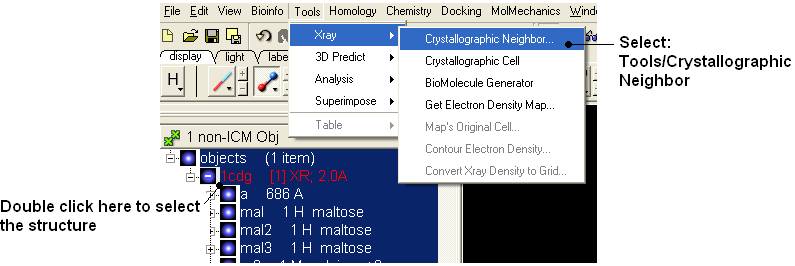
- Tools/Crystallographic Neighbor
- Select a 7A radius
- Check "create symmetry related molecules" and "display symmetry neighbors".
- Inspect the neighbors surrounding maltose(mal). Each symmetry related subunit can be colored by object by clicking and holding the representation button in the display tab and selecting color-by.
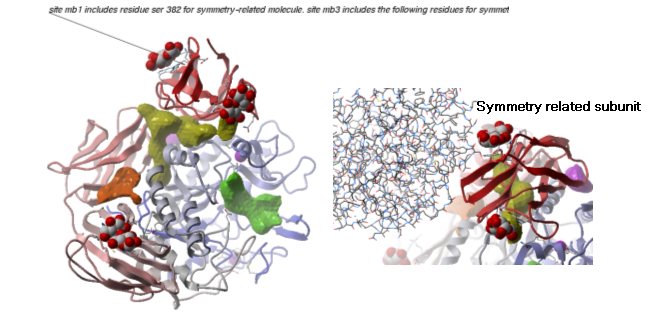
22.8.2 PDB Preparation - Occupancy and B-Factors |
Background When preparing a PDB for analysis (eg docking or modeling) it is important to check the reported occupancies and b-factors. The occupancy is a fraction of atimic density at a given center. If there are two eqally occupied conformers both will have an occupancy of 0.5 - the normal value is 1 range 0-1. The *{B-Factor} is the mean-square displacement of atom from its position in the model - the normal range is 5-50.
One way of visualizing the occupancy and b-factor is by coloring the structure by these values. You can do this by clicking and holding on a representation button in the display panel and selecting Color-by.
As an example let us look at the crystal structure 1ATP
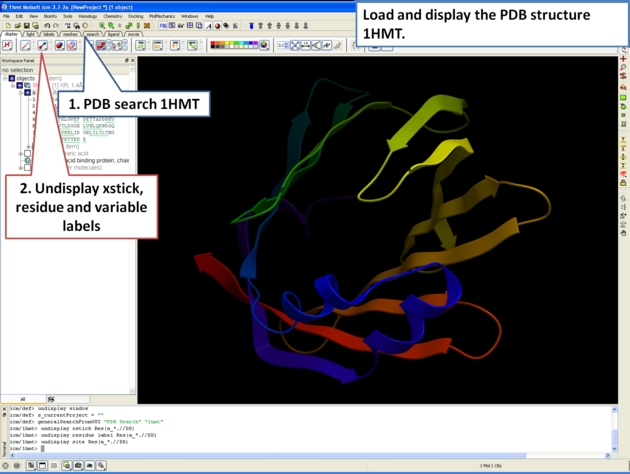
- Type in the PDB search tab 1atp and the structure will be displayed in the graphical display.
- Use the ICM workspace to undisplay everything except for the "e" subunit. You can do this by clicking in the blue boxes in the ICM Workspace.
- Display the "e" subunit in wire representation using the wire button in the display tab.
- Click and hold on the wire button and select Color-by B-Factor. Regions of high B-factor are colored red.
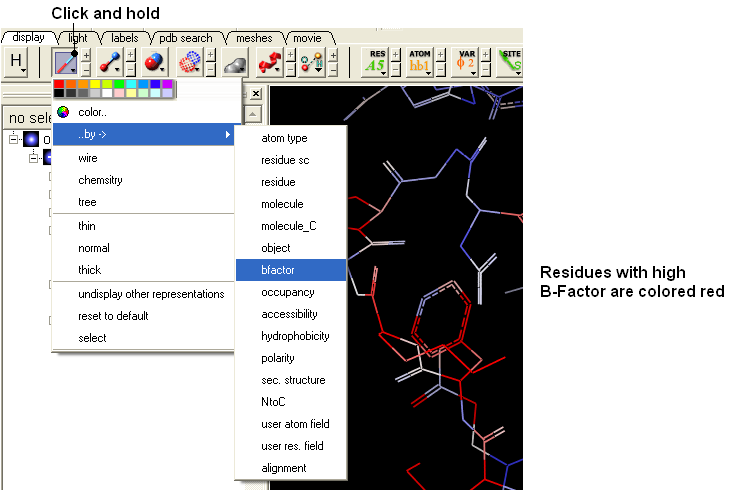
22.8.3 PDB Preparation - Residue Alternative Orientation |
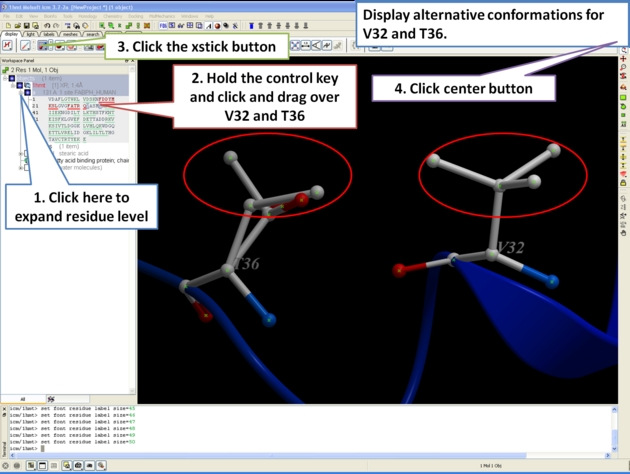
For some very high resolution structures two alternative conformations for a residue are provided. Therefore for docking you need to decide to use one conformation of the residue or generate seveal separate docking models. This could be performed using multiple receptor conformation docking.
Here is an example of alternative residue orientations found in a crystal structure of a Fatty Acid Binding protein in complex with stearic acid.
22.8.4 Biomolecule Generator |
Objective
Here we will investigate the biological environment of a virus protein . PDB code 1DWN.
Background
It is very useful to know how a protein from the PDB may look in a biological environment. The PDB entries solved by X-ray crystallography and deposited in the PDB contain the information about the crystal structure rather than the biologically relevant structure. For example, for a viral capsid only one instance of capsid protein complex will be deposited and only one or two molecules of haemoglobin that is a tetramer in solution maybe deposited. In some other cases the asymmetric unit may contain more than one copy of a biologically monomeric protein. ICM reads the biological unit information and has a tool to generate a biological unit. Not every PDB entry has the biological unit information.
Instructions
- Read and load the PDB file 1DWN
- Tools/Xray/Biomolecule Generator
- Tick the makeAllBiomolecules box (Warning this may take a few minutes to generate)
- The generated molecules will be listed in the ICM Workspace. Each one can be selected and displayed. The biomolecule is shown below.
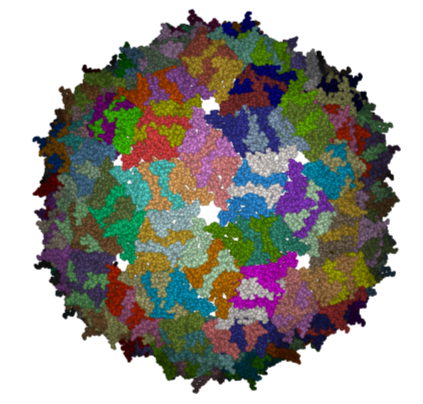
| NOTE: Please note that right clicking on a PDB file in the ICM Workspace will tell you whether there is any Biomolecule information available for the structure. If this information is not present then the option will be greyed out. |
Manual References (Web Links)
| Prev Superimpose | Home Up | Next Tutorial - Working with the Molecular Editor |